In late 2016, I was excited. I had observed (Barnum et. al 2018) that a set of simple communities enriched from bay mud contained abundant bacteria from the Candidate Phyla Radiation (CPR) and an abundant archaeum from the DPANN superphylum. CPR and DPANN organisms may form large portions of the tree of life and share a strange biology: they are obligate epibionts of other cells. The lifestyle of these symbionts may teach us more about microbial biology and ecology, their parasitism of other cells may be exploited as biocontrol agents, and the way they form connections between cells could be translated into biotechnology. And – at that point – had only been isolated in co-culture 5 times. Here were several of these enigmatic organisms, sometimes among the top 10 most abundant genomes in a community.
I thought, what if I could find a way to isolate these organisms in co-culture? Being well-versed in how targeted cultivation can fail from teammates’ experiences in my lab, I wondered if there were ways to do so in an untargeted, scaleable way.
I devised a promising approach, spoke with a professor on campus who was an expert on these organisms, gathered some simple laboratory supplies, and self-assuredly struck out to isolate the CPR and DPANN. Unfortunately, a third-year PhD student with a good idea has no power to achieve it without support from supervisors and funders, and I failed to win that support. I stopped my attempt, returned to my dissertation research on chlorine, and started pushing my one good idea on others at UC Berkeley. (Like a normal grad student, I did so merrily at seminars and grumpily at happy hours). I thought I should share this idea more broadly, but I worried doing so before could have interfered with others’ research.
Now, the field has reached a turning point where many groups have had recent success cultivating these small, epibiotic microorganisms. In particular, Batinovic et. al 2021, discussed below, showed me that “the cat is out of the bag.” I bet that several research groups have co-cultured isolates awaiting publication. I expect that similar ideas have been discussed in groups at conferences. I hope that other groups have considered / are implementing what I will suggest here and will succeed soon. (If you are reading this: don’t worry, no one will scoop you because of this blog post! You got this!). All of this is to say that I don’t view this as an original insight. I am no longer employed at an academic institution but still consider myself part of the community, and I think my community would benefit from having a more open discussion about this interesting cultivation problem.
The Standard Approaches
It is ridiculous to call the successful isolation of a hard-to-isolate organism as “standard,” as it is nothing but herculean or inspired or (as you will see in some cases) flat-out lucky. That being said, these are the approaches that have thus far been successful, in reverse chronological order, to my knowledge:
- Batinovic et. al 2021 (behind the paper) applied filtered wastewater fluid onto plated lawns of Gordonia amarae (phylum Actinobacteria) and whole-genome sequenced clear plaques that formed on those lawns. Many of the plaques were bacteriophage, but one of the plaques was Candidatus Mycosynbacter amalyticus (phylum: Saccharibacteria / fmr. TM7).
- Moreira et. al 2021 (behind the paper) used a micromanipulator attached to a light microscope to physically pick an epibiont, Vampirococcus lugosii (phylum: Absconditabacteria / fmr. SR1), and its host, a Halochromatium-related species (phylum: Proteobacteria), from a lake water culture enriched for anoxygenic phototrophs. They then sequenced the organisms.
- Hamm et. al 2019 enriched and isolated Halorubrum lacusprofundi strain R1S1 in hypersaline water, later detecting a Nanohaloarchaeum in metagenomes of the enrichment culture. Fluorescence activated cell sorting of Ca. Nanohaloarchaeum antarcticus (phylum: Nanohaloarchaeum) from the enrichment allowed for its reliance on its host to be confirmed.
- Cross et. al 2019, rather remarkably, used proteins encoded in TM7 cells (phylum: Saccharibacteria / fmr. TM7) to create antibodies that could selectively capture these organisms and their hosts (various Actinobacteria), which were subsequently plated.
- Golyshin et. al 2017 with sequencing identified Candidatus Mancarchaeum acidiphilum (phylum: Micrarchaeota) in a culture of the acidophile Cuniculiplasma divulgatum (phylum: Thermoplasmatota).
- St. John et. al 2017 used qPCR to monitor enrichments for Nanoarchaea, having a breakthrough when adding 0.22-micrometer filtrate into defined cultures of isolates, upon which they obtained a stable enrichment of Ca. Nanoclepta minutus (phylum: Nanoarchaeota) and its host Zestosphaera tikiterensis (phylum: Crenarchaeota).
- Wurch et. al 2016 also used qPCR to monitor enrichments for Nanoarchaeota, followed by optical tweezer selection of a single host-epibiont pair, a successful hyperthermophilic and acidophilic co-culture of Ca. Nanopusillus acidilobi (phylum: Nanoarchaeota) and its host Acidilobus sp. 7A (phylum: Crenarchaeota).
- He et. al 2015 enriched for Nanosynbacter lyticus TM7x (phylum: Saccharibacteria / fmr. TM7) using antibiotics and only detected it with PCR in plated colonies with Actinomyces odontolyticus XH001 (phylum: Actinobacteria). The connection between cells was apparent with light microscopy. See Utter et. al 2020 (behind the paper) for an important follow-up.
- Huber et. al 2002 observed under a light microscope that a hyperthermophilic “isolate” (nice try!) of Ignicoccus hospitalis (phylum: Crenarchaeota), obtained from a shallow marine hydrothermal vent, was covered by Nanoarchaeum equitans (phylum: Nanoarchaeota).
- Guerrero et al. 1986 originally described Vampirococcus (phylum: Absconditabacteria / fmr. SR1) after observing it with light microscopy attached to Chromatium (phylum: Proteobacteria) cells in environmental samples, which inspired Moreira et. al to carry out their study, but its special evolutionary position would not be recognized until this year.
By my count, these incredible successes have resulted in the following number of species:
- CPR Bacteria: 5 Saccharibacteria and 2 Absconditabacteria.
- DPANN Archaea: 3 Nanoarchaeota and 1 Nanohaloarchaeum.
The isolations fall into several categories:
- Serendipitous: observation of an epibiont with an already isolated host (e.g. Huber et. al).
- Casual: identification of an epibiont with a host, and a subsequent pain-free isolation (e.g. Moreira et. al).
- Targeted: purposeful co-cultivation of an epibiont and host, often with many attempts (e.g. St. John et. al).
- Targeted, made easier by expensive investment: development of a method to selectively pull an epibiont and host into isolation (e.g. Cross et. al).
Should others try to reproduce these methods? If researchers take the extra step to look for epibionts while isolating other strains, “casual” isolations should increase. And by all means, go have a “serendipitous” discovery. Unfortunately, all of the “targeted” methods share a big drawback: intense effort focused on a single host or epibiont. We should admire these researchers’ effort and success while asking:
Is there a better way?
A Potential Untargeted, Scaleable Approach
Consider Batinovic et. al 2021 from a different perspective. From a habitat filled with many different epibiotic species and many host species, they (1) isolated a potential host, (2) selected all potential epibiotic species (with a 0.2-micrometer filter), (3) re-introduced those potential epibiotic species to the isolated potential host species, (4) watched for changes in the isolate that indicated the colonization of the isolate by an epibiont, and (5) confirmed the colonization of the epibiont.
You will, I expect, immediately ask yourself a series of questions:
What if you repeated the process but with a different potential host?
What if you repeated the process but with 100 different potential hosts?
What if you repeated the process but with epibionts collected from many samples from the same habitat?
How many more hosts and epibionts could you find?
The sum answer to these questions it that there is no reason to think that by scaling the isolation process and not focusing efforts on a single targeted host or epibiont, you would dramatically increase the likelihood that you obtain epibionts in co-culture.
Let’s consider there is a probability that a isolatable bacterium or archaeum could be a host of a epibiont present in its habitat (P_host), a probability that a epibiont among a sample could be cultivated (P_epibiont), and a probability that the colonization of the host by the epibiont can be observed (P_observation). What is the probability that you can get the epibiont in co-culture (P_get_epibiont_for host)? Perhaps a reasonable set of assumptions – that hosts are relatively rare, that a epibiont for that host won’t always be present in a community, and that an assay for observation would miss half of cases – the probabilities would be P_host = 0.1, P_epibiont = 0.2, and P_observation = 0.5. The joint probability would be P_get_epibiont_for host = P_host * P_epibiont * P_observation = 0.1 * 0.2 * 0.5 = 0.01. So, you have a 1 in 100 chance of getting a epibiont for a single potential host. But if you try 100 potential hosts, according to the binomial distribution, your probability of a 0 success is P_failure = (100!/ 0! (100-0)!) * (P_get_epibiont_for)^0 * (1-P_get_epibiont_for)^(100-0) = 0.37, or about 2 in 3 chance of success. Not bad! For 1000 potential hosts, with a naive assumption of the probabilities being independent, the probability of 0 successes is miniscule (0.0001).
My thinking on how to do this has not changed much over the years. I believe that these steps will work but encourage you to reevaluate them:
- Choose an appropriate habitat.
- Isolate many potential hosts.
- Select potential epibionts by cell size, removing viruses and free-living cells as much as possible, and concentrate.
- Grow host cultures with and without the addition of the epibiont concentrate.
- Assay for the growth of epibionts with the host. Confirm with microscopy.
- Isolate specific host-epibiont pairs.
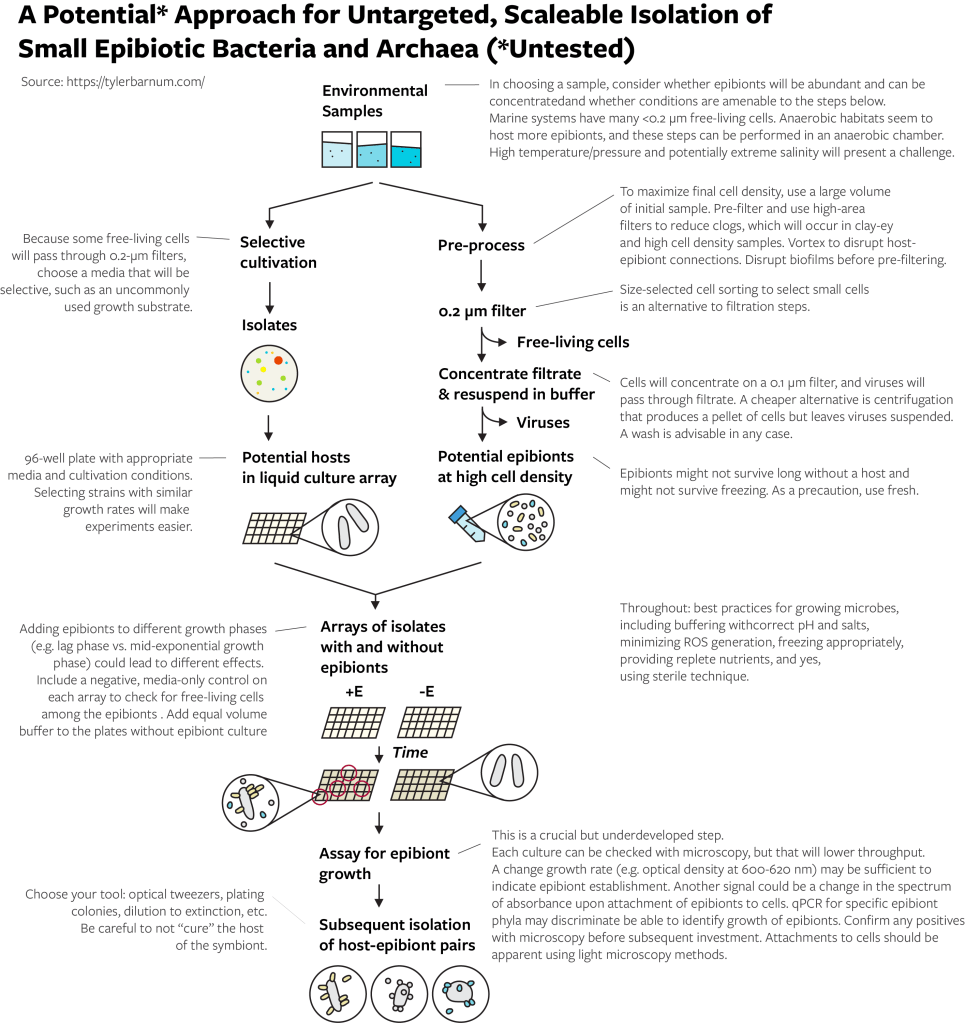
How you implement these steps depends on your available equipment and your ability to increase throughput. The below method flowchart provides some suggestions and important notes. I played around with this enough in 2017 (very little) to know that the size selection and assay for epibiont growth are important steps to get right. For example, you need to use a filtration set up that will not get clogged by large volumes of liquid. For another example, if your assay is microscopy, looking at a microscope over and over again will take a lot of time. For yet another example, contrary to common opinion, free-living cells can and will pass through 0.2-micrometer filters (especially certain marine bacteria), so you should think about ways to limit their growth with a selective growth medium. Equipment to sort cells by size is not widely available but may be a huge benefit for the initial collecting of epibionts and for finding and collecting host cells to which epibionts are attached.
A note on habitat and host strains. Everyone has a bias towards their familiar habitat or strain. I beg you, before you choose the same study system you have always worked with, consider how this will limit your likelihood of success. Have the presence of these organisms ever been demonstrated in your habitat of study? Do your host strains resemble wild strains, or have they been propagated in the laboratory for years? Please, be critical and set yourself up for success. As a default, I would suggest a highly connected habitat with high cell densities, fewer environmental filters, and moderate diversity. For example, saturated sediments from wetlands or streams would be a better place to start than agricultural soils or animal microbiomes. I know little about the biology of epibiotic microorganisms, but I presume they benefit from transport between hosts (hence highly connected, fewer filters) and a high availability of hosts (high cell density). Moderate diversity would seem to decrease the chance that a habitat has no cultivable host-epibiont pairs (i.e. low diversity) and the chance that the epibiont for an isolated host is at low relative abundance (i.e. high diversity). Avoid a community if you know that it only has one or two epibiont phylotypes because if there are only one or two hosts, you will likely not succeed in isolating either. If you have PCR primers for CPR or DPANN phylum of interest, testing a sample for a diverse set of these organisms using amplicon sequencing would be a smart check on your choice before additional work.
Why Has This Not Been Done Before?*
I don’t know.
I believe the description of some specific host-epibiont relationships has led to a feeling that host-epibiont pairs are rare and must be carefully teased out of a community. You must identify the host, with the epibiont, then isolate both. Some of the above papers make it clear that this is not the case in cultivation, or that an epibiont can be hosted by species from different genera. I do not argue that some or many epibionts can be specific to one host species. But assuming that even only a small fraction of epibionts have a very broad host range provides enough justification to pursue an untargeted approach by increasing P_epibiont in the above math.
I believe that microbiologists could more readily adopt higher-throughput approaches but may be constrained by what’s affordable or what’s familiar. (In some cases, the affordability concern continues the issue of valuing non-human resources over the time and output of very smart, talented, driven students and staff). Additionally, microbiologists who study epibionts may have invested time in learning metagenomics and be uncomfortable with investing more time learning cultivation methods, resulting in fewer researchers in this subject area working on cultivation.
Finally, I believe that these is a notion that uncultivated organisms must be fastidious, or hard to cultivate. In many cases, an organism may truly be fragile in cultivation conditions or reluctant to grow quickly. One can imagine how epibiotic bacteria and archaea could be slow to form attachments to their host, fail to have a growth rate equaling a fast growing host, etc. However, several of the above publications show that at sufficient cell densities, some epibionts can be removed from their host, filtered, and transferred to a different host, successfully! There is no reason to think these organisms are not robust and cultivable just because they haven’t been cultivated in large numbers. Instead of comparing them to uncultivated free-living cells, a more apt comparison would be to consider them like uncultivated phages: happy to proliferate when provided a suitable host.
So, while this cultivation strategy has not been done, I believe that it will work and provide easier access to study a unique type of biology.
* This phrasing was a bit of misdirection. Take note that the question should really be, “why has this not been done before at scale?” Batinovic et. al 2021 cultivated a Saccharibacteria organism with basically this approach. I am simply suggesting to do what they did, with more potential hosts and an approach tuned away from phages and towards epibiotic cells.
Happy cultivating, and good luck!
How should I cite this article?
You probably should not cite this article because it is a blog post. I did no publishable work, but if your research benefitted from this article, I would be pleased to see a mention in your acknowledgements section. I would most enjoy hearing how this helped you find something cool. I do not have any time to help because I spent it all on writing a blog post.
